DAPT
Welcome to the official documentation of Descriptor-based Adaptive Perturbation Transformer (DAPT)!
📍 Overview & Motivation
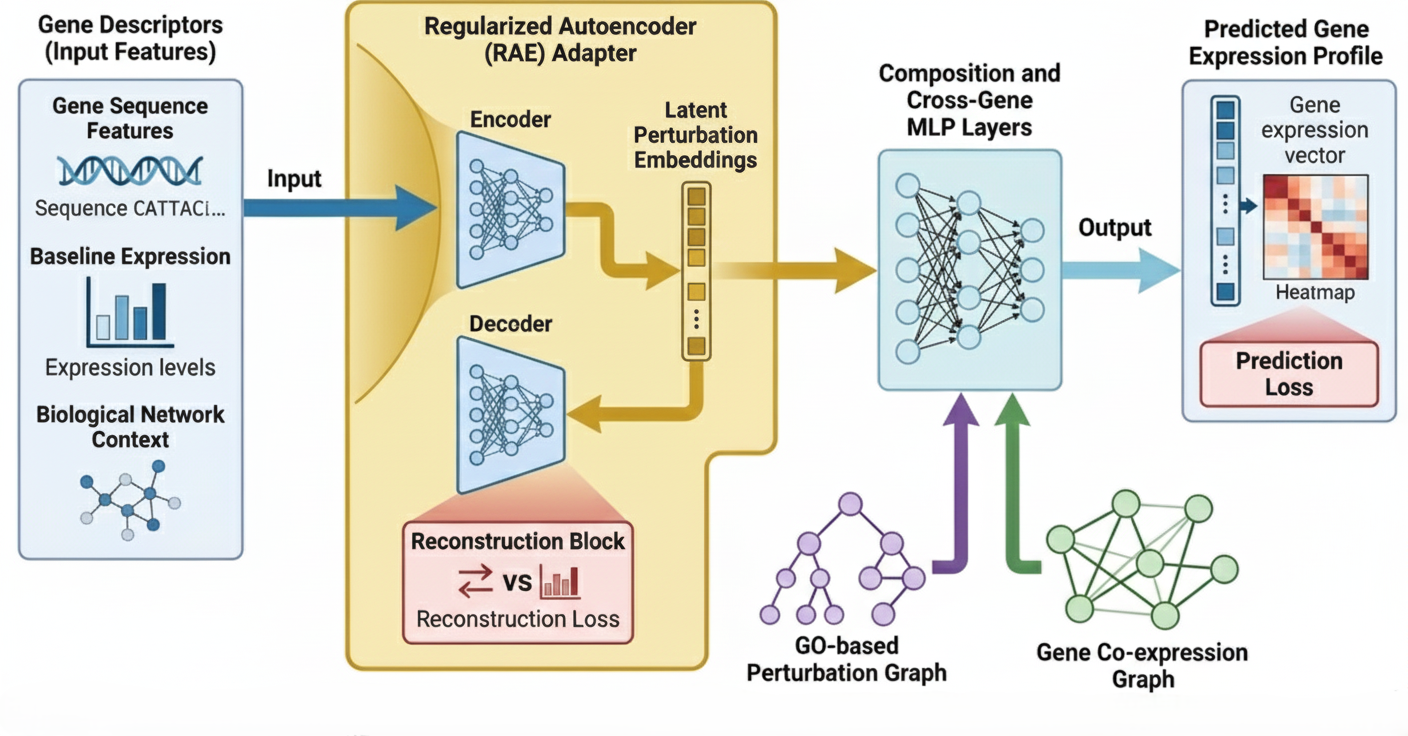
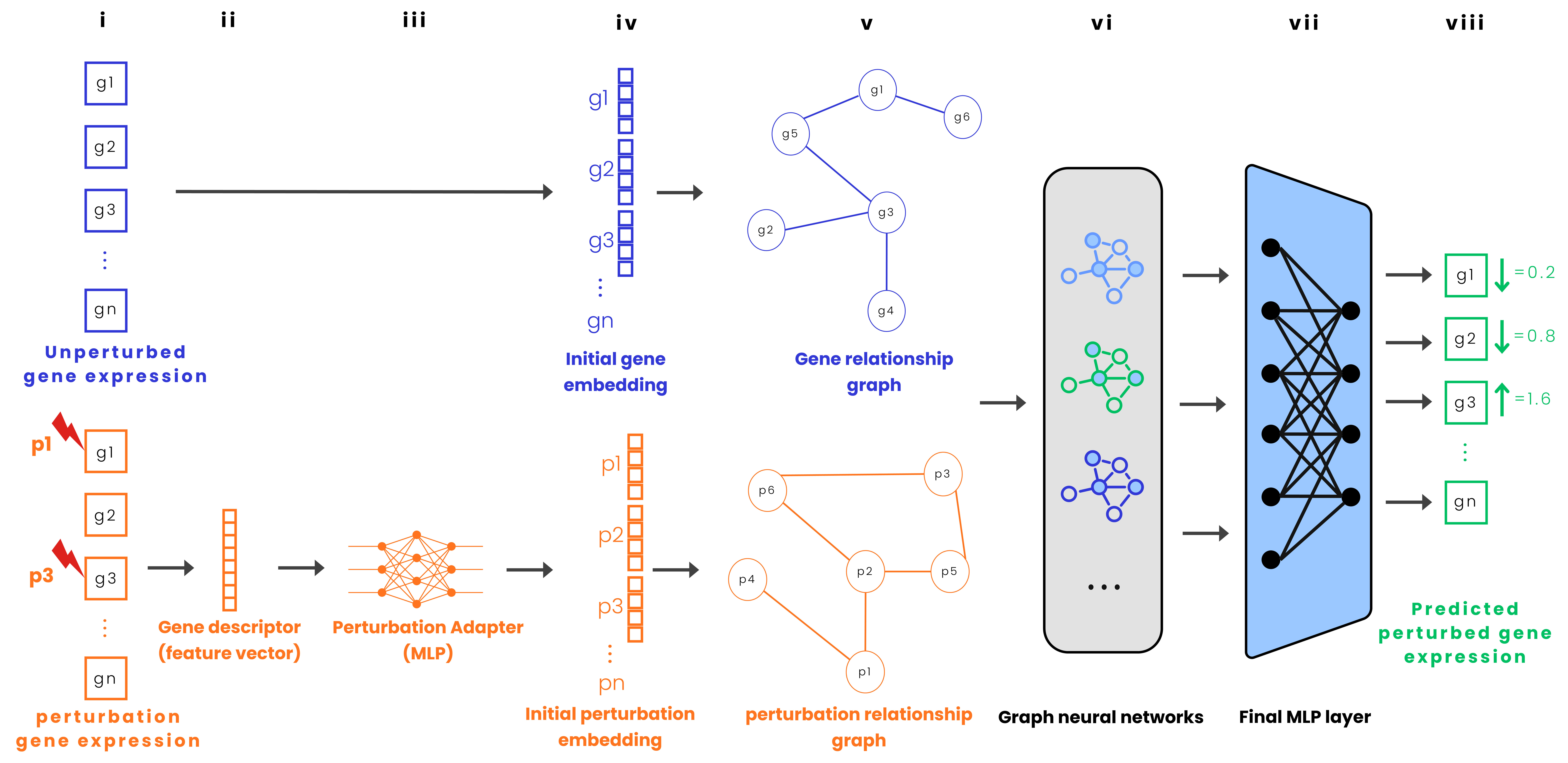
Understanding how gene perturbations alter cellular transcriptional responses is central to modern biomedicine, enabling rational therapeutic design and the identification of genetic interactions. While recent single-cell CRISPR screening platforms like Perturb-seq provide large-scale datasets, existing computational methods exhibit instability when predicting multigene perturbations, particularly for genes unseen during training or entirely missing from biological knowledge graphs.
We present DAPT, a novel deep learning framework designed to predict the transcriptional outcomes of single and multigene perturbations. Developed to overcome the structural limits of existing models. DAPT introduces a descriptor-driven perturbation adapter integrated with Graph Neural Networks (GNNs) and prior biological knowledge.
DAPT replaces standard ID-based perturbation embeddings with a descriptor-based representation. This allows our model to learn the intrinsic biological properties of genes, significantly improving zero-shot prediction for both unseen and out-of-vocabulary (OOV) perturbations.
💡 Key Innovations
- Descriptor-Based Representation: Replaces discrete, ID-based embeddings with semantic biological gene descriptors, enabling parameter sharing across genes.
- Perturbation Regularized Autoencoder (RAE): Transforms high-dimensional descriptors into dense latent embeddings through non-linear transformations.
- Biological Knowledge Integration: Utilizes Gene Ontology (GO) and Gene Co-expression graphs as dense substrates for message passing, ensuring predictions are grounded in functional pathways.
- OOV Generalization : Generates perturbation representations directly from input descriptors, enabling out-of-distribution generalization, particularly for unseen or novel gene perturbations.
🚀 Quick Start
DAPT is designed to be flexible. You can interact with it visually through a web dashboard, or integrate it directly into your own bioinformatics scripts for high-throughput batch processing.
1. 🔗 Interactive Web Dashboard
The easiest way to experience DAPT is through our interactive web application—no installation required! For real-time inference and visual analysis of single and multigene perturbations in Norman et al. (2019) Perturb-seq dataset.
👉 Access the DAPT Web Dashboard Here
How to use the dashboard?
1. Select a Task: Use the left sidebar to choose your experiment mode.
2. Choose Target Genes: Select the specific genes you want to perturb from the dropdown menus.
3. Run Inference: Click the "Run DAPT Inference 🚀" button to process the Co-expression and GO Graphs.
4. Explore Results: Open the "📊 Top Genes Chart" and "📋 Full Data Table" tab to visualize the most shifted genes plot and complete predicted transcriptomic profile.
2. ⚙️ Programmatic Python API
For researchers wanting to evaluate large datasets, DAPT provides a clean API.
Here is a minimal example of how to load the data manager (Constellation), initialize the Dapt neural network controller, and predict a multigene perturbation:
import torch
from dapt import Constellation, Dapt
# 1. Initialize the Data Manager
# Constellation loads the biological mappings, GO graphs, and datasets
constellation = Constellation(config_path="configs/norman_experiment.yaml")
# 2. Initialize the DAPT Controller
model = Dapt(config=constellation.get_model_config())
model.load_model("weights/dapt_best_weights.pt")
# 3. Predict Post-Perturbation Expression
# Example: Predicting a complex non-additive double-gene perturbation
baseline_expression = constellation.get_baseline()
predicted_transcriptome = model.predict(
unperturbed_state=baseline_expression,
perturbation_set=["A1BG", "CD86"]
)
print(f"Predicted Expression Profile Shape: {predicted_transcriptome.shape}")
Developed by NG Ka Ho as a Final Year Project at City University of Hong Kong (Department of Computer Science).